Page path:
- Organic Geochemistry
- Research
- Deep biosphere
Hinrichs Lab - The deep subseafloor biosphere
Introduction
Jule Verne’s famous book A Journey to the Centre of the Earth reports how Professor Lidenbrock, his nephew Axel and their guide Hans meet a versatile realm of living organisms on their descend into the interior of our planet. When the novel was first published in 1864, the idea of life existing in the deep subsurface remote from sunlight and air was genuine science fiction. But towards the end of the 20th century, when scientific drilling opened the window into hundreds of meter deep environments, it became clear that Jules Verne in a sense was right. Deep under the terrestrial ground and the seafloor life is very abundant (Whitman et al., 1998; Parkes et al., 2000; Pedersen, 2000), albeit mainly in the form of prokaryotic microorganisms which are not visible to the naked eye. In fact, the deep subsurface biosphere represents a substantial fraction of Earth’s total biomass; first estimates suggest a contribution of 10 - 30% (Parkes et al., 1994; Whitman et al., 1998).
The discovery of the deep biosphere gives rise to new, fundamental questions about the evolution and distribution of life. Considering the broad range of temperatures and high pressures in the subsurface and the lack of fresh organic matter as a source of energy and nutrients, the existence and survival of an extensive deep biosphere are remarkable. Geofuels, i.e. gases that are generated by mineral-water interactions or derived from the deep crust, might be an alternative energy source in the absence of light and photosynthesis. So far, only little is known about the individual inhabitants of the deep biosphere, about their genetic diversity, metabolic capabilities and survival strategies. Moreover, the relationships between deep subsurface ecosystems and the surface world remain to be explored.
Deep biosphere research has been a rapidly growing field during the last decade. The Hinrichs Lab contributes to this field with a focus on the exploration of life in the deep subseafloor.
Deep biosphere research has been a rapidly growing field during the last decade. The Hinrichs Lab contributes to this field with a focus on the exploration of life in the deep subseafloor.
Access to deep subseafloor samples: scientific ocean drilling
For the exploration of the deep biosphere access to unadulterated samples is a key challenge. In the case of the deep subseafloor, samples for scientific research started to become available with the Deep Sea Drilling Project (DSDP) in 1968 and studies of pore-water chemistry soon suggested the existence of metabolically active microorganisms in deeply buried sediments. Microbiological investigations, however, were only initiated in the subsequent Ocean Drilling Program (ODP) towards the end of the 1980s when the acridine orange dye method and rigorous contamination tests enabled reliable direct counts of total cell numbers in drill cores. Systematic investigations conducted by a group of scientists around Prof. John Parkes (University of Bristol, UK) revealed that even 1 km below the seafloor 1 milliliter of sediment often still contains more than 1 million intact cells. Following the discovery of abundant microorganisms in deeply buried sediments ODP Leg 201 was devoted to the exploration of life in the deep subseafloor in 2002 and the deep biosphere became a major research initiative in 2004 when ODP was succeeded by the Integrated Ocean Drilling Program (IODP), an international effort involving 24 countries lead by implementing organizations in the U.S., Japan and Europe. Cores retrieved by DSDP, ODP and IODP are stored in three central repositories, one of which is located in the MARUM building on the campus of the University of Bremen (Bremen Core Repository, BCR).
Our research approach
The investigation of life deep below the seafloor is a challenging task, not only because the retrieval of samples is elaborate and expensive but also because the very low biological activity in this ecosystem limits the applicability of cultivation based microbiological and biogeochemical methods.
Without a doubt successful cultivation of single deep biosphere species is an important goal, yet alternative methods are needed to achieve a representative characterization of the deep biosphere. Both molecular biology and organic geochemistry provide such methods that do not require growth of microorganisms in the laboratory but extract information from biological molecules that are produced in situ in the natural environment. Molecular biology targets genetic information that is encoded in DNA and RNA molecules. The genetic information allows microbiologists to distinguish individual species and to gain insight into the building plans for their metabolism. While in a sense molecular biology sheds light on the blue print of life in the deep biosphere, organic geochemistry investigates the building blocks that are actually used.
In the Hinrichs lab, we are particularly interested in membrane forming lipids, in substrates, products and intermediates of microbial metabolism, and in the processes that link the deep biosphere to global biogeochemical cycles.
Without a doubt successful cultivation of single deep biosphere species is an important goal, yet alternative methods are needed to achieve a representative characterization of the deep biosphere. Both molecular biology and organic geochemistry provide such methods that do not require growth of microorganisms in the laboratory but extract information from biological molecules that are produced in situ in the natural environment. Molecular biology targets genetic information that is encoded in DNA and RNA molecules. The genetic information allows microbiologists to distinguish individual species and to gain insight into the building plans for their metabolism. While in a sense molecular biology sheds light on the blue print of life in the deep biosphere, organic geochemistry investigates the building blocks that are actually used.
In the Hinrichs lab, we are particularly interested in membrane forming lipids, in substrates, products and intermediates of microbial metabolism, and in the processes that link the deep biosphere to global biogeochemical cycles.
Intact polar lipids in cell membranes provide information about the inhabitants of the deep biosphere
Polar lipids in cell membranes are fragile molecules that rapidly decompose after cell death. Therefore, we can use intact polar lipids (IPLs) as biomarkers that contain information about life cells (for more information see “Biomarkers"). Unlike the genetic code, the composition of cell membranes is not specific for individual species, but still Bacteria and Archaea produce distinctly different IPLs that can be used to identify single clades of these two domains of life. We use qualitative and quantitative analyses of IPLs to determine the composition of microbial communities living in the deep biosphere.
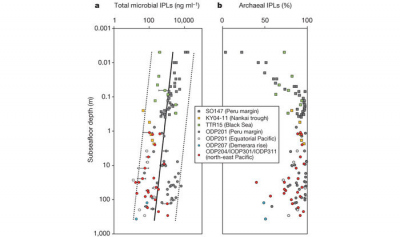
a, Concentration of total IPLs versus depth. Regression lines show total microbial IPLs and 95% prediction interval (dotted lines). b, Relative contribution of archaeal IPLs to total microbial IPLs. (Figure taken from Lipp et al., 2008)
Our systematic investigation of IPLs in the upper 360 meters of sediment in various regions of the world ocean has revealed that Archaea are not only widespread in the deep subseafloor but also much more abundant than Bacteria (Lipp et al., 2008; Lipp & Hinrichs, 2009). Complementary molecular biological investigations allocated these Archaea almost exclusively to phylogenetic lineages, that have only recently been discovered in deep subseafloor environments and never been cultivated in the laboratory (Lipp et al., 2008).
Though Bacteria are often barely detectable in situ, we found concentrations of bacterial IPLs to increase remarkably during storage of samples under oxic conditions at +4°C while at the same time concentrations of archaeal IPLs remained virtually constant (Lin et al., 2010). The latter underlines the importance of appropriate sample treatment and storage.
Archaea are a poorly understood domain of life. They have long been considered to be exotics that only occur in extreme environments like hot springs and salt lakes. Clearly, this view is now changing with the discovery of archaeal dominance in the deep subseafloor also suggesting a significant contribution of Archaea to the global biomass.
Though Bacteria are often barely detectable in situ, we found concentrations of bacterial IPLs to increase remarkably during storage of samples under oxic conditions at +4°C while at the same time concentrations of archaeal IPLs remained virtually constant (Lin et al., 2010). The latter underlines the importance of appropriate sample treatment and storage.
Archaea are a poorly understood domain of life. They have long been considered to be exotics that only occur in extreme environments like hot springs and salt lakes. Clearly, this view is now changing with the discovery of archaeal dominance in the deep subseafloor also suggesting a significant contribution of Archaea to the global biomass.
Following the discovery of Archaea in the deep subseafloor, benthic archaea have become a major research endeavor in the Hinrichs Lab. With DARCLIFE we have initiated a project to study their carbon cycle, life strategies, and role in sedimentary ecosystems with an interdisciplinary team of scientists. This project is funded through an Advanced Grant of the European Research Council (for more information see “DARCLIFE”).
Organic metabolites in pore-waters reflect biogeochemical processes in the deep biosphere
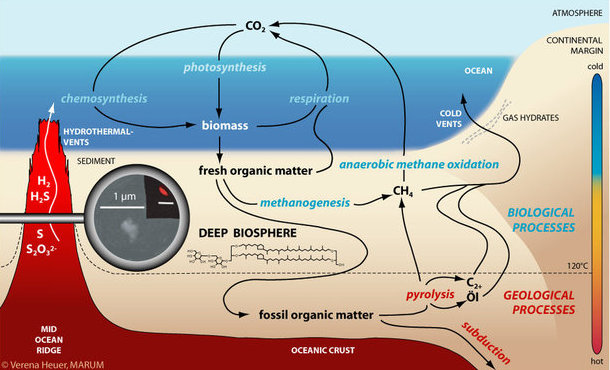
Hydrocarbon gases: methane, ethane, propane
Methanogenesis and anaerobic oxidation of methane (AOM) are key processes in the cycling of carbon in the seafloor. To our current knowledge, both processes can only be mediated by Archaea. They link the deep biosphere to climate via the flux of methane into the water column and atmosphere.
Since AOM controls the flow of methane out of the sediment, this process has been studied intensively at cold seeps where methane reaches the seafloor. On a global scale, however, AOM in deeply buried sediments might play a similar or even larger role in the regulation of atmospheric methane contents and therefore calls for a better understanding of the involved processes and organisms.
We use the natural abundance of stable carbon isotopes to investigate the fate of methane (for more information see "biogeochemical processes"). Since the light carbon-12 isotope is strongly preferred during methanogenesis, methane from biogenic sources has a distinct isotopic composition and its consumption by AOM and uptake into biomass can be tracked by stable carbon isotope analyses. While in shallow sediments at cold seeps stable carbon isotope analysis show the uptake of methane carbon into the membrane lipids of methanotrophic Archaea (Elvert et al., 1999; Hinrichs et al., 1999), there is no indication for the uptake of methane carbon into archaeal cells in deep AOM zones even though AOM accounts for a major fraction of carbon cycled in these ecosystems (Biddle et al., 2006). Moreover, we also found carbon isotopic evidence for the in situ microbial production of ethane (ethanogenesis) and propane (propanogenesis) in cold subseafloor sediments (Hinrichs et al., 2006).
Since AOM controls the flow of methane out of the sediment, this process has been studied intensively at cold seeps where methane reaches the seafloor. On a global scale, however, AOM in deeply buried sediments might play a similar or even larger role in the regulation of atmospheric methane contents and therefore calls for a better understanding of the involved processes and organisms.
We use the natural abundance of stable carbon isotopes to investigate the fate of methane (for more information see "biogeochemical processes"). Since the light carbon-12 isotope is strongly preferred during methanogenesis, methane from biogenic sources has a distinct isotopic composition and its consumption by AOM and uptake into biomass can be tracked by stable carbon isotope analyses. While in shallow sediments at cold seeps stable carbon isotope analysis show the uptake of methane carbon into the membrane lipids of methanotrophic Archaea (Elvert et al., 1999; Hinrichs et al., 1999), there is no indication for the uptake of methane carbon into archaeal cells in deep AOM zones even though AOM accounts for a major fraction of carbon cycled in these ecosystems (Biddle et al., 2006). Moreover, we also found carbon isotopic evidence for the in situ microbial production of ethane (ethanogenesis) and propane (propanogenesis) in cold subseafloor sediments (Hinrichs et al., 2006).
Volatile fatty acids: acetate
We develop new methods for stable carbon isotope analysis of organic metabolites that we find dissolved in pore-waters (Heuer et al., 2006) (see also "metabolites"). Our main interest is directed towards volatile fatty acids because they are central intermediates in the degradation of organic matter and important substrates for microorganisms. Acetate is particularly interesting since it is not only formed by fermentation and thermal alteration of organic matter during sediment burial (Wellsbury et al., 1997) but also by reduction of carbon dioxide with hydrogen (acetogenesis). Thus, it could potentially transform geofuels, i.e. gases that are generated by mineral-water interactions, into an organic substrate that can be used by a broad range of microorganisms (Pederson, 2000).
Our investigations in methane rich sediments at the northern Cascadia Margin (NE Pacific, IODP Site 1329) reveal a large variability in the stable carbon isotopic composition of pore-water acetate and do not only point to an active and metabolically diverse deep biosphere but also suggest an important role of acetogenesis (Heuer et al., 2009).
We combine stable carbon isotope analysis of organic metabolites with thermodynamic calculations and molecular biological investigations to learn more about the biogeochemical processes and the organisms that mediate them. On the northern Juan de Fuca Ridge Flank (NE Pacific, IODP Site 1301), where we found stable carbon isotopic indications for on-going acetogenesis, complementary functional gene analysis were conducted by our collaborators and confirm the presence of a wide range of aerobic and anaerobic (sulfate-reducing) Proteobacteria, Gram-positive sulfate-reducing Bacteria, and uncultured branches of Bacteria or Archaea with the ability to perform acetogenesis (Lever et al., 2010).
Our investigations in methane rich sediments at the northern Cascadia Margin (NE Pacific, IODP Site 1329) reveal a large variability in the stable carbon isotopic composition of pore-water acetate and do not only point to an active and metabolically diverse deep biosphere but also suggest an important role of acetogenesis (Heuer et al., 2009).
We combine stable carbon isotope analysis of organic metabolites with thermodynamic calculations and molecular biological investigations to learn more about the biogeochemical processes and the organisms that mediate them. On the northern Juan de Fuca Ridge Flank (NE Pacific, IODP Site 1301), where we found stable carbon isotopic indications for on-going acetogenesis, complementary functional gene analysis were conducted by our collaborators and confirm the presence of a wide range of aerobic and anaerobic (sulfate-reducing) Proteobacteria, Gram-positive sulfate-reducing Bacteria, and uncultured branches of Bacteria or Archaea with the ability to perform acetogenesis (Lever et al., 2010).
These findings point to different metabolic processes in deep and shallow sediments.
Outlook
With our on-going research we are aiming to gain a deeper understanding of life in the subseafloor at the interface between biological and geological processes. We are determining the composition and metabolic activities of subseafloor ecosystems. We are investigating the energy and nutrient sources of these ecosystems and the impact of environmental properties on the composition of microbial communities. We are aiming to understand the role of the deep biosphere in the global carbon cycle, including its dependence on biological processes in the surface world, its interactions with geological processes in sediments and oceanic crust, and its effects on the chemistry of the ocean and atmosphere.


